Data extraction from a data warehouse at a German university hospital – data quality and current challenges
Dorothee A. Busch 1,2Mareike Przysucha 1
Jens Hüsers 3
Jonathan M. Mang 4
Cornelia Erfurt-Berge 2
Ursula Hübner 1
1 Research Center for Health and Social Informatics, Osnabrück University of AS, Osnabrück, Germany
2 Department of Dermatology, Uniklinikum Erlangen, Friedrich-Alexander-Universität Erlangen-Nürnberg (FAU), Erlangen, Germany
3 Hamburg, Germany
4 Medical Center for Information and Communication Technology, Uniklinikum Erlangen, Friedrich-Alexander-Universität Erlangen-Nürnberg (FAU), Erlangen, Germany
Abstract
Treatment of chronic wounds is a topic with high research potential, however, routine data are hardly standardized and cannot be accessed easily. The introduction of data integration centers within the Medical Informatics Initiative (MII) in Germany aims at making routine care data available for research. In this study, we seek to investigate the degree of structuredness and the quality of clinical data for clinical research on chronic wounds. These data were extracted in csv format from the data warehouse in the data integration center at the University Hospital in Erlangen. We found that data retrieved for wound research were heterogeneous in format (free text vs. structured data) and quality (missing values, inconsistencies, lack of longitudinal data, missing context) and were mainly based on billing relevant data. Basic epidemiological and clinical questions might be answered, but necessary data are missing for deeper analyses of treatment outcomes. A more clinically driven and standardized documentation would improve the availability of data relevant for research on the courses of treatments in the field of wound care and beyond.
Keywords
data quality, wound care, EHR, observational studies, data integration center
Introduction
Data gathering in clinical research on chronic wounds
Chronic wounds are a major burden for both the individual patients and the society, causing a significant reduction in quality of life and accounting for high annual costs [1]. Evidence-based management is essential to improve patient outcomes. To better understand the patient population with chronic wounds and the treatment response to different therapies, it is helpful to evaluate routine data, ideally beyond a single center. In addition to randomized controlled trials and other controlled studies, routine data offer a promising method to create evidence, especially within areas such as wound care, where limited studies are available [2], [3]. However, routine data is rarely documented in a standardized, structured way, which makes evaluations difficult.
There are comprehensive recommendations for information to be documented, for leg ulcers [4], [5], but a uniform definition of what information should be included in wound documentation in daily practice is currently not available. Additionally, wound related data are currently stored in different formats, applications, and with varying content [6].
This heterogeneity of data makes a comparison of information and the export of data for further analysis a difficult task. In addition, patients are treated by different specialties, such as dermatology, plastic surgery, or vascular surgery due to the interdisciplinary nature of the challenges that have to be addressed for stabilizing or healing the wounds.
One approach to obtain data sets of an appropriate size in a reasonable time is to make routine data accessible to health professionals and health data scientists in a FAIR manner [7]. In Germany, this idea led to the foundation of the Medical Informatics Initiative (MII) in 2016, with four consortia consisting in total of all German University Hospitals [8], [9]. Each participating hospital set up a data integration center (Datenintegrationszentrum, DIC). Each DIC extracts data from IT systems within the hospital, harmonizes them, stores them in a standardized way, and makes them accessible for local and cross-site research following a process that enables (a) exploring the feasibility of the study and (b) warranting data protection according to regulatory requirements. The structure of the data at the end of the transformation process is defined in the MII core data set [10], consisting of base modules (e.g., patient, encounter, diagnoses, procedures, laboratory data, and medication) [11] and extension modules like oncology or microbiology data [12], [13]. After obtaining an ethical vote, a positive vote of a Use and Access Committee (UAC) and if necessary, anonymizing the data, healthcare providers, but also other research institutes are allowed to analyze these data. The main export format of the data is HL7® FHIR® R4 [14], but formats like comma separated value (CSV) and also formats of data in an earlier step in the transformation process may be exported.
Data from a DIC can be analyzed within the data holding facility or, after an UAC voting and export, at the institution that requested the data. For cooperation with external partners for data analysis, there are two ways to ensure data privacy: either the data are covered by the broad consent [15], [16] or they are anonymized, e.g. by removing or masking identifying information, or by giving ranges containing the original data, possibly resulting in an anonymization bias [17].
Current way of data gathering at the University Hospital Erlangen
The work presented here is a case study from the University Hospital Erlangen which belongs to the MIRACUM consortium of the MII [8], [9]. At the Department of Dermatology, as in many other medical institutions, manual chart review of data is still the most frequently used method for data retrieval to answer clinical and scientific questions. A local Data Warehouse (DWH) incorporated into the DIC, is used to identify all patients for a given time period based on the ICD-10-GM [18] diagnoses provided. Subsequently, all patient records of the included patients are manually searched, the necessary information is manually transcribed or copied and sorted to be used for the analysis. In addition to this effort, the heterogeneity of the data is a challenge for the analysis. Therefore, data for clinical research and its quality is often limited [19], leading to a reduction of the strength of evidence.
The DIC at the University Hospital Erlangen
In Erlangen, the local DWH contains a copy of data from almost all digital systems, including the hospital’s enterprise resource planning system (SAP) and the clinical documentation systems recording patient information from routine clinical processes. For local analyses (e.g., controlling, individual evaluations, local research), the extraction of a data set can be requested via the DIC following regulated access processes shown in Figure 1 [Fig. 1]. A study plan with a scientific question and a positive ethical vote as well as a data usage application must be provided and consent from the heads of all data-producing departments must be obtained. A cost estimate will be provided for data extraction based on the effort and scope of the request and must be approved by the institution requesting the data. The interdisciplinary UAC then assesses the application. If the assessment is positive, the request is forwarded to the DIC. If data are intended to be shared with an external project partner, they must be anonymized unless there is a legal basis for sharing, such as a study-specific consent or a broad consent.
Figure 1: Process to obtain data from the DIC in Erlangen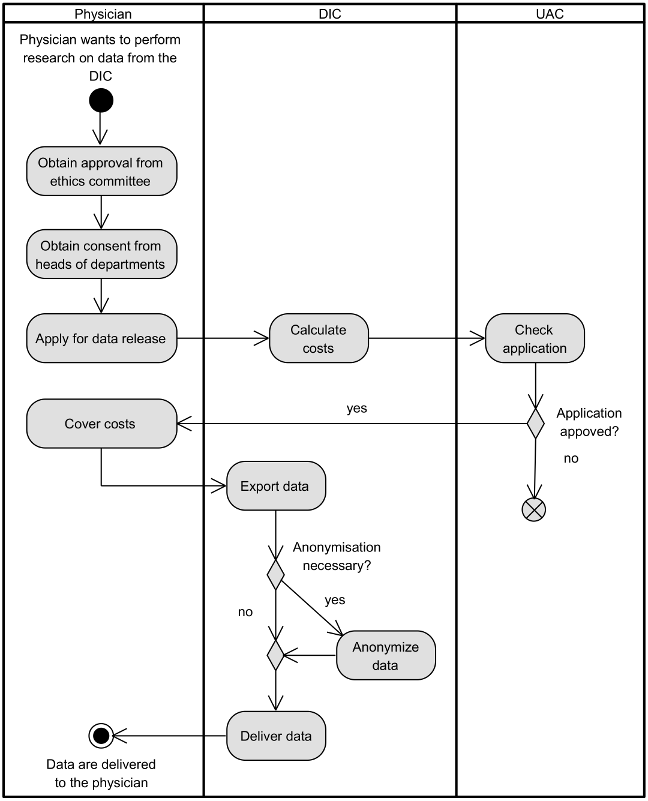
Objective
To our knowledge, there has been no use case involving DIC data in the field of wound care to date. The overall objective of this study is therefore to examine the structure and quality of data extracted from the data warehouse of the data integration center at Erlangen University Hospital. This data is based on the MII core data set. Specifically, the following questions about the data will be answered:
- How is the data quality in terms of completeness, relevance, and accuracy?
- Can the data that are based on the MII core data set be used to answer relevant clinical and epidemiological research questions based on the core dataset?
Furthermore, this study seeks to get insights into the impact of an anonymization process when sharing data with external academic research partners. This study served as a pilot for evaluating the datasets for wound care obtained through the German Research Data Portal Healthcare (FDPG) and represents preliminary work for a larger follow-up study to address these questions on a broader scale.
Methods
Data gathering
We applied for data from the DIC at the University Hospital Erlangen and followed the process shown above. Our data analysis was guided by typical clinical and epidemiological questions, specifically looking at the composition of the patient group, the treatments and their outcomes, the use of medication, and comorbidities, like:
- How can the patient cohort be described in terms of age, gender, wound diagnosis, primary diagnosis, comorbidities, and underlying conditions?
- What treatments and procedures were applied, what medication was given, and what are the outcomes in terms of laboratory parameters, wound parameters, and condition/problems?
- How do these interventions and outcomes change over time?
As data should originate from the field of wound care, we requested data for a cohort of patients with chronic wounds (ICD-10-GM codes L97, I83.0, I83.2, I87.01, I87.21, I70.24, L88, E10.75) being treated at the Dermatology Department from the period January 1, 2021 to March 30, 2023, leading to data until March 29, 2023.
Assuming that the clinical research questions mentioned above, which guided the exploration for data structuredness and quality, could be answered by the MII core data set we requested these data. Next to data of the base modules [11], we asked for data of the extension module microbiology [13]. Wound images were not included in this study as they are not part of the MII core data set.
Therefore, consent for data usage was required from the heads of the Central Laboratory and Microbiology Department. A consent to obtain clinical data from the Dermatology Department was not necessary, as the Dermatology Department itself filed the request. After calculating costs and obtaining the necessary consent from the different heads of departments, an application for data extraction was filed.
The data were exported from the data warehouse into six csv files, each one containing one of the base modules (patient, encounter, diagnosis, procedure, medication, laboratory values). As the extracted data were forwarded to the University of Applied Sciences Osnabrück where we planned to analyze them, they needed to undergo k-anonymization [20] with k=5 due to German General Data Protection Regulations.
Data analysis
To get first insights into the data provided, we created a Unified Modeling Language (UML) class diagram of the data provided. Data types were empirically distilled from the available data. Then we determined whether data were coded and whether these codes came from a local, national or international code system. We also looked at the distribution of the data and whether data were anonymized.
Afterwards, we performed a data quality analysis on the data, considering the factors completeness, relevance, and accuracy.
Completeness means, that “all values for a certain variable are recorded” [21]. Therefore, the number of missing values were determined.
Relevance generally is defined as the “extent to which data are applicable and helpful for the task at hand” [22]. In our case, a physician (DB) rated the clinical relevance of the data by whether the data could potentially be used to answer clinical and epidemiological research questions. For laboratory data and antibiograms, the data were grouped by the identifying code, and first results were listed for each code. A physician rated all codes for their general clinical and epidemiological relevance.
Accuracy is understood as the measure to express whether “data are correct, reliable and certified free of error” [22]. As the original data and exported data cannot be compared in this study, the accuracy within our study focused on laboratory data. Stated data types were tested, and whenever different representations of the same result within one entry were provided, we tested whether these representations were congruent. If string and numeric values differed, a physician determined the most plausible value. Additionally, the consistent spelling of units was verified.
Results
The information model for the data provided is given in Figure 2 [Fig. 2]. Due to data anonymization, the original dates for all clinically relevant data were exchanged with the difference in days to the date of the first encounter. As reference, the age at the first encounter and the timeframe of this encounter was given for each patient.
Figure 2: Information model of the data provided to the University of Applied Sciences Osnabrück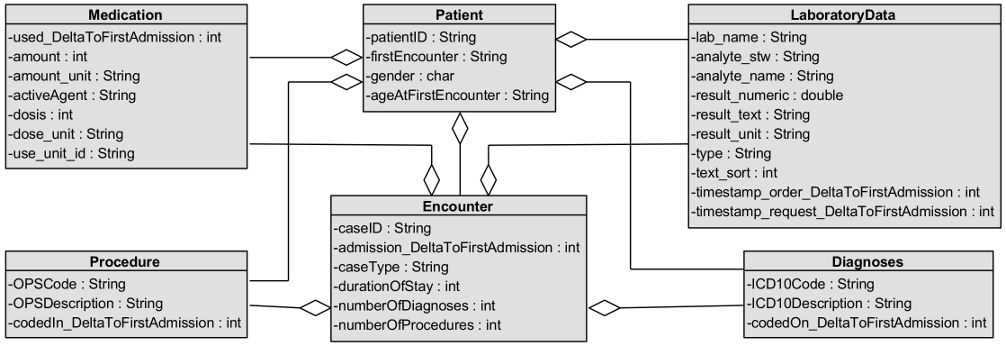
We received data on 499 patients and 1,186 encounters, comprising a total of 4,775 diagnosis entries, 1,555 procedure entries, 201 medication entries, and 143,349 laboratory entries, as shown in Table 1 [Tab. 1]. Except for medication, relevant data were available in a structured way.
Table 1: Overview of data obtained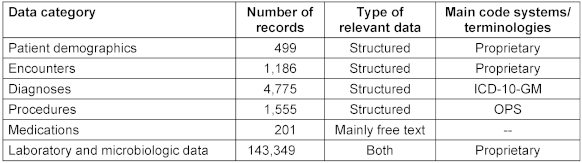
Table 2 [Tab. 2] shows the number of missing data of all data categories within the data provided. Each patient record included all data on patient demographics. The same held true for the diagnoses and procedures data.
Table 2: Frequencies of missing data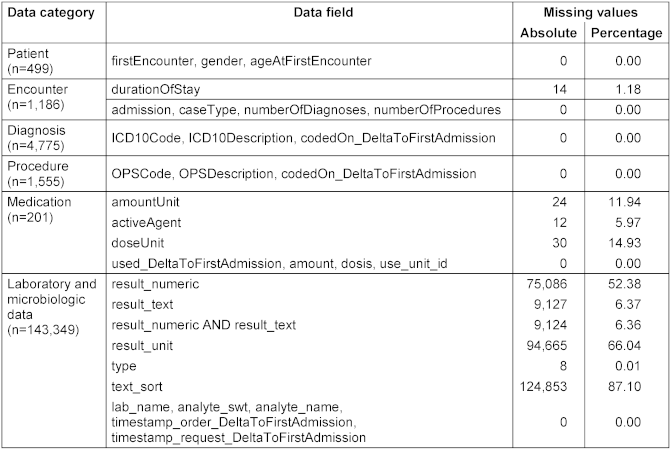
Within the patient demographics, gender was coded using a local code system with the codes W (weiblich = female), M (männlich = male), and * (no description). The information on the age at first encounter was clustered with partially large ranges, e.g., 19 to 97 years. The change of gender codes to the code * as well as the clustering of the age resulted from the anonymization procedure.
Encounter data were nearly complete. Only 14 out of 1,186 encounters had no duration of stay. Among the encounter data, the admission dates and times were replaced by the month or year of the first encounter, e.g., ‘May 2022’ or ‘2023’ as a result of the anonymization. All remaining dates were then given as a difference in days to the first encounter.
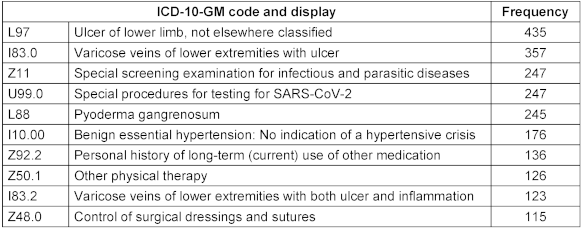
Regarding diagnoses, the ICD-10-GM codes used for billing (using SAP software) were provided. In Table 3 [Tab. 3] the ten most frequent diagnoses are shown. Four of them are wound diagnoses (L97, I83.0, L88, I83.2), two are specific codes for SARS-CoV-2 screening before admission (Z11, U99.0), two are typical codes for comorbidities (I10.00, Z92.2) and two are codes for general and wound specific treatments (Z50.1, Z48.0). In the data provided, there were no signs for anonymization.
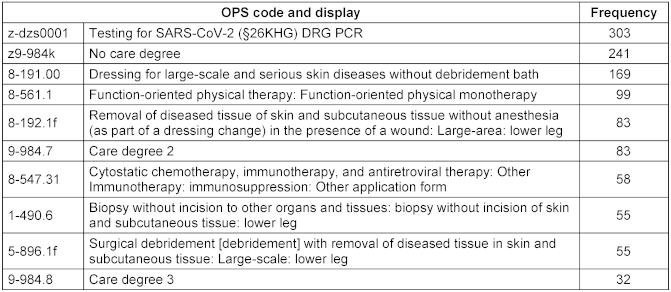
The Procedure data provided were coded with the national system of operations and procedures (OPS) [23], extended by a few local codes. The ten most frequent procedures are shown in Table 4 [Tab. 4]. Notably, testing for SARS-CoV-2 was the most frequent procedure. The care degree (either as no degree or the degree number) was also frequently documented. Four procedures (8-191.00, 8-192.1f, 1-490.6, 5-896.1f) referred to interventions concerning the wound. Similar to diagnoses, mostly billable information was captured. In total, procedures were only captured for less than two thirds of the patients (309 out of 499, 61.9%). Alike the diagnosis information, there were no signs for anonymization visible.
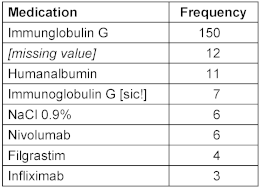
Medications were not stored as structured data, only the active ingredients had been recorded as free text. The eight most frequently documented medications are shown in Table 5 [Tab. 5], all other medications had a frequency of 1. Only for 21 out of 499 patients (4.2%) and 43 out of 1,186 encounters (3.6%), medication data were available. In 12 out of the 201 medication records, the active_agent was missing. Whether this was due to the anonymization process or because of the data in the DWH could not be determined. Also, the dataset did not include the patients’ long-term medication, but predominantly billing-relevant medication, mostly intravenously applied medication, such as immunotherapies. Medications that are taken orally were not included.
The laboratory values and antibiograms of the microbiology, coming in total from 14 different laboratories, were provided comprehensively. As described in Figure 2 [Fig. 2], these data contained, among other information, a local code to identify the content (analyte_swt) and a human readable description for the code (analyte_name). The identifying codes came from a local code system but could have been also provided as LOINC codes by the DIC. Out of 1,776 different codes, 1,567 (88.2%) were rated as relevant, covering 129,761 (90.5 %) of all 143,349 entries, the remaining 209 codes (11.8%) were rated as non-relevant, covering 13,588 of all entries (9.5%). Examples for relevant codes include codes for leucocytes, hemoglobin, erythrocytes, and hematocrit, while among the non-relevant codes were hints or dates.
For reporting a laboratory value, two rows with the names result_numeric and result_text were provided. As can be seen in Table 2 [Tab. 2], the column result_numeric held information in 68,263 entries, result_text in 134,222 entries, and at least one of the columns were present in 134,225 entries. Figure 3 [Fig. 3] shows the distribution of result_numeric and result_text. In total, 9,124 entries contained no result at all. Among these, units were provided only for 297 entries (3.3%). While most units indicated numeric values (e.g., %, x 10^3/ul, g/dl)), two units did not reveal the data type (–, Material), covering six entries. In the remaining 8,827 entries, the unit was missing, making it nearly impossible to determine the real data type.
Figure 3: Distribution of result_numeric, result_text, result_unit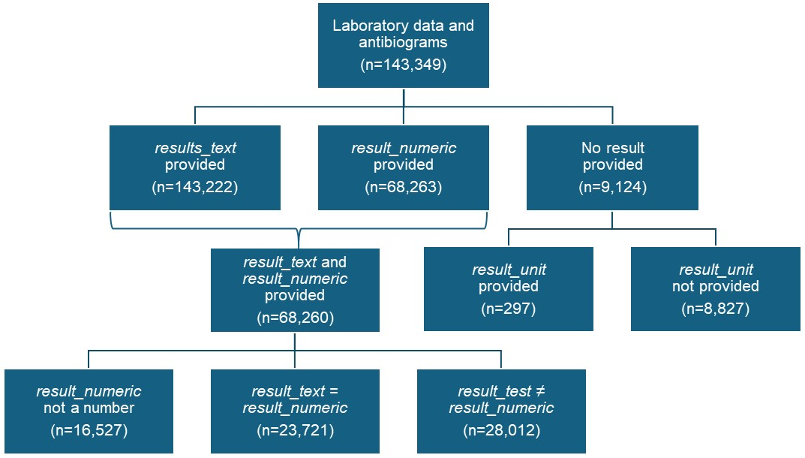
Therefore, in total, 65,962 entries contained textual results, 68,554 entries contained assumably numerical results, and 8,833 entries contained data of unknown type as neither a result nor a unit were stated.
Among the 134,225 entries with a result (string and/or numeric representation), 68,260 entries contained a string as well as a numeric representation. When type-casting the numeric content to a number (it was stored as a string before), 16,527 entries in the column result_numeric could not be transferred into a number (also see Figure 4 [Fig. 4]). For the remaining 51,733 data entries the type-cast was successful. A comparison of both columns result_numeric and result_text showed that the columns corresponded to each other in 23,721 entries, while in 28,012 entries, the string and the numeric value differed. The analysis of the multiplication factor between the two values showed that both results differed by factor 10 (2,388 entries), by 100 (140 entries), by 1,000 (5 cases) and by ≈109 (25,479 entries). For these 28,012 laboratory values, the textual representation was more likely to be the correct data than the numeric value, e.g., Calcium 2.06 mmol/l (text + unit) compared to 20,599,999,428 mmol/l (numeric value + unit).
Figure 4: Distribution of correctness of numeric laboratory results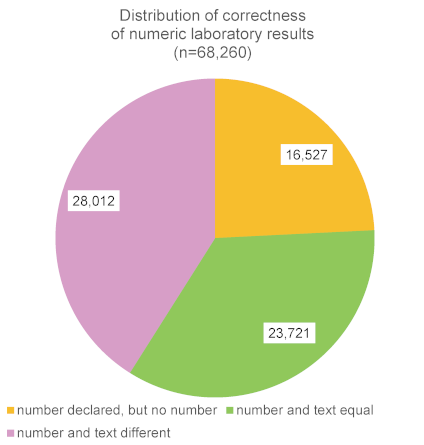
As described above, 13,588 of the laboratory data contained no valuable information, while 129,761 data were relevant in principle. The distribution of the relevant data by data type is shown in Table 6 [Tab. 6]. Among the relevant values, numeric data had a higher proportion to be relevant (67,804 out of 68,554, 98.9%) in comparison to textual data (55,822 out of 68,263, 84.6%).
Table 6: Distribution of relevance by data type of laboratory data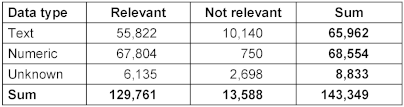
If applicable, units were also given, but the spelling of the same unit differed slightly between the different laboratories, e.g., for erythrocytes, the units “x10^6/µl“ (791 entries), “x 10^6/µl” (106 entries), and “x 10^6/ul” (602 entries) were used.
Discussion
General remarks
After asking for data from the MII core data set, we received data from a prefinal transformation step of data delivery of the core dataset, leaving some of the harmonization and standardization steps undone. To our knowledge the reasons were that (a) we also requested images, which are not part of this study, and the paths to these images were not yet available in HL7® FHIR®, and (b) the data had to be anonymized for Osnabrück and the anonymization routine only worked on table-based data. The following discussion will address issues in terms of heterogeneity of the format (free text vs. structured data) and the quality criteria completeness, relevance, and accuracy, particularly in terms of missing values, inconsistencies, lack of longitudinal data and missing context. The question whether there were data missing that had been recorded elsewhere cannot be answered.
Missing values
The data contained a relatively high number of missing values especially for laboratory values and antibiograms. Possible reasons for missing data might be a) data missing in the source system, b) data deleted/not extracted during the transformation process from the original source to the DWH, or c) data deleted during the anonymization process ensuring k-anonymity (k=5). The reasons could not be determined in this study. Therefore, the assessment of the completeness might partly underrate the number of existing values considering the data available at both the hospital information system or the data warehouse. However, incomplete data limit the relevance of the data and their capability to render the complete picture. Missing values produced during the anonymization process hint at the limited opportunities of external partners to make full use of the data.
Inconsistencies and data format
Data inconsistencies were mainly observed for units in the laboratory data, specifically as different systems used different representations of the same units. This leads to an increased workload for data analysts and can increase the likelihood of misinterpretation of the data. By using international standards like UCUM [24] for units this problem might be overcome. Other issues are free text formatted laboratory values and problems with data which are either not numeric though it was stated differently, or faulty regarding the decimal separator.
The MII core data set requires data to be represented in the HL7® FHIR® format, incorporating standardizations and harmonizing steps, which can be delivered by the DIC. For laboratory values, the main FHIR® resource is the Observation Resource [14]. The definition of the laboratory observation from the MII [25] defines that UCUM codes must be used for units, leading to a uniform machine-readable representation. It also allows IT systems to distinguish the data type into quantities with given separators, coded concepts with the corresponding code systems, and free text, but also allows the system to mark information as missing either due to unavailability or due to masking. Extracting data at a later transformation step and implementing the data structure as defined in the MII core data set may therefore enhance the accuracy of the data and allow a better assessment of the completeness of the data. It can be expected that a better understanding of the data becomes possible, and thus an easier analysis of the structured data is ensured. However, the challenges posed by free text data would remain.
Relevance and completeness
SAP as source of the data
A large part of the data was extracted from the hospital’s enterprise resource planning system (SAP system), which is used to code for billing purposes with ICD-10-GM codes. These codes provided important clinical information to determine the distribution of diagnoses in the population and to make statements about common comorbidities. However, it is important to note that, according to the knowledge of the first author of this study (DB), especially diagnoses and procedures relevant to billing had been recorded and the complete list of clinical diagnoses and procedures was not available. In other words, the extracted data were based on DRG-relevant diagnoses and procedures, rather than on medical relevance. This means that certain aspects, such as the main diagnoses (as ICD-10-GM code), and procedures such as wound debridement, were well-documented and easily accessible, while other important underlying illnesses, such as former amputation, organ insufficiencies, depression, or reduced compliance, were rarely extracted.
At the University Hospital Erlangen, diagnoses for medical treatment are recorded in another information system (“Soarian”) for outpatient treatment, often using free text. These diagnoses were not extracted by the DIC, leaving out many comorbidities. Also, information about the patients’ long-term medications and most of the medications and procedures during the hospital stay was not available, although this information was stored in one of the digital documentation systems. This information, once extracted, could be useful to get a better overview of the patients’ treatment. The MII core data set allows data of different modules to be exported as long as they can be extracted in a structured way. It does not restrict data to be only billing relevant or need to originate from the hospital itself. The inclusion of structured data from the national electronic health record may improve the availability of structured data which do not originate from the hospital within the hospital information system and may therefore improve the availability of data within the DIC.
The strong focus of information on being billing-relevant, also holds true for medication and procedures. Therefore, drawing conclusions about treatment response based on the laboratory values over time (e.g., decreasing inflammatory blood markers) is speculative and cannot be linked to individual treatments as not all information is accessible. As a consequence, the most interesting clinical questions, namely those about treatment outcomes, cannot be answered by the data provided. We could not include more data as they were not available in a meaningful amount before 2021. However, with the inclusion of more longitudinal data and from more specialties originating from further data sources particularly clinical information systems, this problem might be overcome, and further data might become available.
Lack of contextual information
The data obtained also lacked some contextual information. For each encounter the number of procedures and diagnoses were given, but the information provided did not contain information about the main diagnosis. This leads to a potential bias in the dataset as also patients with a wound who presented for other reasons (main diagnosis) were included. Due to the fact that all encounters considered included patients of the department of dermatology only, it is fair to assume that the patients came to get their wound treated. However, this assumption may not be valid for all patients.
In the HL7® FHIR® representation, it is possible to represent most of these contextual data. The representation of an encounter in an Encounter resource allows the additional indication of meta-data, e.g., department main diagnoses or admission diagnosis [26].
Relevance and completeness – reorganization of information
Among the laboratory data considered irrelevant there was the request date of the laboratory analysis and there were also comments and grouping codes. In HL7® FHIR®, it is possible to distinguish between a ServiceRequest resource to represent the request for a laboratory analysis of a specimen, including a request date, and the actual results. Observation resources contain the actual laboratory value, possibly with a free text comment, which are summarized in DiagnosticReport resources which also allow conclusions to be given [25]. Therefore, a representation in HL7® FHIR® according to the core data set definition by the MII would help organizing information in a meaningful way and allows researchers to focus on relevant information.
Anonymization
A significant portion of the data was unavailable due to anonymization. In line with the principles of k-anonymization (k=5), extremely broad age ranges were assigned in some instances, rendering the data unsuitable for further analysis, but also information was masked, like gender, or probably also deleted. Other options would have been (a) to analyze the data at the University Hospital Erlangen in accordance with recent regulations in Bavaria, or (b) to ask for data for which a broad consent was given [15], [16]. As option (a) – analysis of data at the University Hospital Erlangen by one of their staff members (DB) – is a suitable possibility for the authors for future analyses of the data, option (b) – analyze data for which a broad consent was given – was not taken into account, because of an extremely limited number of patients who had given a broad consent. The amount of data accessible under the broad consent will be only a fraction of the anonymized data. In future studies researchers need to decide whether they would prefer all data with lower quality due to anonymization steps or more complete data of less patients. It should also be considered that the Health Data Utilization Act (Gesundheitsdatennutzungsgesetz, GDNG) of 2024 allows health care providers to share health related patient data under certain conditions (Section 6 (3) sentence 4 of the GDNG). As the request for the data was filed before that date and as the external academic partner in this study is not a health care provider, the GDNG still would not have allowed data sharing without data anonymization (Section 6 (3) sentence 3 of the GDNG). Also, first experiences with the GDNG in terms of sharing data with other health care providers in a non-anonymized way, which is allowed under certain circumstances (Section 6 (3) sentence 3 of the GDNG), are still pending. Thus, studying the impact of anonymization on data expressiveness is still a relevant case.
Analysis of wound care related data
This study inspected and gauged data very similar to the MII core data set for its usability to answer epidemiological and treatment related clinical questions in wound care. Although wound related diagnostic, laboratory and treatment data were available in principle, their meaningfulness was limited as discussed above. This renders any clinical analysis difficult, like questions on the impact of an antibiotic therapy on infection markers, or the treatment of multi-resistant germs. Epidemiological studies based on the anonymized data are partly possible, but impossible in other ways. The distribution of age, patients per month, living area of patients (via zip code), insurance status, and partly gender among different groups of patients is impossible to derive as age, dates and partially gender were masked or transformed. Research questions like the distribution of wound types and number of wounds per patient, distribution of nursing degree, readmissions, and type of encounter and the identification of multi-resistant germs were possible to answer while analyses on comorbidities, procedures applied, and medications prescribed or administered were limited as the relevant data were not yet extracted. Ways to improve the usability of the data include, among others, the inclusion of other data sources, but also a structured documentation allowing an exact extraction of data for research, as the automatic code annotation of free text is prone to errors or misses [27].
Whether these main results also hold true for other conditions not represented as an MII use case like wound care, should be evaluated in future studies. In any case, the development of more use cases within the MII is desirable to obtain more meaningful data across different clinical research areas. In the future, the development of the use case “chronic wounds” could incorporate wound related data such as parameters for wound descriptions, e.g., wound size, wound bed, description of peri wound and wound edge, exudate, debris, and odor [4], [5], [28], into the DICs and make these data available for research purposes. A use case “chronic wounds” would not only be interesting from the perspective of different medical specialties, e.g. dermatology, surgery, diabetology, angiology, but also from the viewpoint of nursing. These developments could be coordinated with other initiatives around standardization in wound care, like the development of a national wound summary [28], and make standardized wound related data available for care as well as for research.
In the near future, the Health Research Portal Germany (Forschungsdatenportal Gesundheit – FDPG) will enable researchers to directly access data from several DICs across Germany, providing new opportunities to conduct research using routine clinical data on a national scale. Currently, initial studies are underway to gather experiences with the FDPG. Osnabrück University of Applied Sciences has been selected as one of the first institutions to evaluate and document experiences with the portal. The present study thus serves as a pilot project aimed at assessing the data quality and scope of information available from one of the participating DICs, with the goal of informing future, effective use of the FDPG in health research.
Conclusion
In conclusion, the data available from the data warehouse at the DIC in Erlangen showed a high variability in structuredness and quality. One of the major drawbacks was that they mainly represented billing-relevant diagnoses, procedures, and medications that do not necessarily meet the requirements for medical research. The accuracy, especially of laboratory data and antibiograms may be improved for numeric values as well as for units. In principle, the data were sufficient in quality for relevant analyses of patient demographics, encounters and diagnoses – with the restriction of not knowing the main diagnosis. Drawbacks resulted from the anonymization process. Similarly, the medication information provided was in principle relevant. However, due to the discrepancy between data provided and data potentially available, results drawn from the medication data provided would have a limited meaning. Alike, a significant number of laboratory data were not relevant. Taking all these results into account, clinically relevant research questions will only partially be answerable. Epidemiologic analyses are partly possible due to the anonymization process but will become more meaningful when data are analyzed within the University Hospital. However, reliable statements about the treatment and thus the treatment response will be impossible to date.
Thus, although major advances in data availability took place, medically adequate data for epidemiological and treatment outcome studies are still partially missing. This deficiency makes clinical analyses difficult not only in wound care – be it performed within the hospital or through external academic partners.
A more clinically driven and standardized documentation as well as standardized application programming interfaces would improve the availability of data relevant for research on the courses of treatments in the field of wound care.
Notes
Ethical votes
- Ethics Commission of Osnabrück University AS, vote-no: HSOS/2021/1/5, 05.01.2022
- Ethics Commission of Friedrich-Alexander-University Erlangen-Nürnberg, vote-no: 22-4-Bn, 28.01.2022
Contributions of the authors
- DB was involved in preparing the ethics application for FAU Erlangen-Nürnberg, obtaining the permissions from the heads of departments, elaborating the research questions, and writing the manuscript, and acted as main contact for the DIC in Erlangen.
- MP was involved in planning the study, preparing the ethics application for Osnabrück University AS, performing the analyses on which the experience is based and writing the manuscript, she also acted as connection between IT professionals and clinicians.
- JH planned the study and prepared the ethics application for Osnabrück University AS, was the contact person for all organizational questions, was involved in a first visual quality check on the data, and revised the manuscript.
- JM was the main contact person at the DIC in Erlangen, supported the application for data, exported and anonymized the data, and revised the manuscript.
- CEB supported and supervised DB, participated in the elaboration of the research questions, and revised the manuscript.
- UH supported and supervised MP and JH and drafted and revised the manuscript substantially.
Acknowledgement
This project (ZIEL) was funded by the German Federal Ministry of Education and Research (BMBF) (grant: 16SV8616).
Competing interests
The authors declare that they have no competing interests.
References
[1] Olsson M, Järbrink K, Divakar U, Bajpai R, Upton Z, Schmidtchen A, Car J. The humanistic and economic burden of chronic wounds: A systematic review. Wound Repair Regen. 2019 Jan;27(1):114-125. DOI: 10.1111/wrr.12683[2] Faraoni D, Schaefer ST. Randomized controlled trials vs. observational studies: why not just live together? BMC Anesthesiol. 2016 Oct 21;16(1):102. DOI: 10.1186/s12871-016-0265-3
[3] Röhrig B, du Prel JB, Wachtlin D, Blettner M. Types of study in medical research: part 3 of a series on evaluation of scientific publications. Dtsch Arztebl Int. 2009 Apr;106(15):262-268. DOI: 10.3238/arztebl.2009.0262
[4] Herberger K, Heyer K, Protz K, Mayer A, Dissemond J, Debus S, Wild T, Schmitt J, Augustin M; Konsensusgruppe. Nationaler Konsensus zur Wunddokumentation beim Ulcus cruris: Teil 2: Routineversorgung – Klassifikation der Variablenausprägungen [German national consensus on wound documentation of leg ulcer: Part 2: Routine care – classification of variable characteristics]. Hautarzt. 2017 Nov;68(11):896-911. DOI: 10.1007/s00105-017-4012-6
[5] Heyer K, Herberger K, Protz K, Mayer A, Dissemond J, Debus S, Augustin M; Konsensusgruppe. Nationaler Konsensus zu Wunddokumentation beim Ulcus cruris: Teil 1: Routineversorgung – „Standard-Dataset“ und „Minimum-Dataset“ [German national consensus on wound documentation of leg ulcer: Part 1: Routine care – standard dataset and minimum dataset]. Hautarzt. 2017 Sep;68(9):740-745. DOI: 10.1007/s00105-017-4011-7
[6] Hübner U, Krämer K, Milde S, Thye J, Egbert N. Szenarien zur Bewertung von elektronischen Wunddokumentationssystemen: Die Studie des AOK Bundesverbandes. Wund Management. 2016;10(4):188-195.
[7] GO FAIR. FAIR Principles. 2018 [cited 2025 Feb 20]. Available from: https://www.go-fair.org/fair-principles/
[8] Semler SC, Wissing F, Heyder R. German Medical Informatics Initiative. Methods Inf Med. 2018 Jul;57(S 01):e50-e56. DOI: 10.3414/ME18-03-0003
[9] Knaup P, Deserno T, Prokosch HU, Sax U. Implementation of a National Framework to Promote Health Data Sharing. Yearb Med Inform. 2018;27(01):302-304. DOI: 10.1055/s-0038-1641210
[10] Ammon D, Kurscheidt M, Buckow K, Kirsten T, Löbe M, Meineke F, Prasser F, Saß J, Sax U, Stäubert S, Thun S, Wettstein R, Wiedekopf JP, Wodke JAH, Boeker M, Ganslandt T. Arbeitsgruppe Interoperabilität: Kerndatensatz und Informationssysteme für Integration und Austausch von Daten in der Medizininformatik-Initiative [Interoperability Working Group: core dataset and information systems for data integration and data exchange in the Medical Informatics Initiative]. Bundesgesundheitsblatt Gesundheitsforschung Gesundheitsschutz. 2024 Jun;67(6):656-667. DOI: 10.1007/s00103-024-03888-4
[11] Medizininformatik-Initiative. Basismodule des Kerndatensatzes der MII. [cited 2025 Feb 20]. Available from: https://www.medizininformatik-initiative.de/de/basismodule-des-kerndatensatzes-der-mii
[12] Medizininformatik-Initiative. Erweiterungsmodule des Kerndatensatzes der MII. [cited 2025 Jul 23]. Available from: https://www.medizininformatik-initiative.de/de/erweiterungsmodule-des-kerndatensatzes-der-mii
[13] MII IG Mikrobiologie DE v2025. 2024 [cited 2025 Jul 21]. Available from: https://www.medizininformatik-initiative.de/Kerndatensatz/KDS_Mikrobiologie_V2025/MIIIGModulMikrobiologie.html
[14] HL7® FHIR® Release 4. 2019 [cited 2025 Feb 20]. Available from: http://hl7.org/fhir/R4/
[15] Zenker S, Strech D, Ihrig K, Jahns R, Müller G, Schickhardt C, Schmidt G, Speer R, Winkler E, von Kielmansegg SG, Drepper J. Data protection-compliant broad consent for secondary use of health care data and human biosamples for (bio)medical research: Towards a new German national standard. J Biomed Inform. 2022 Jul;131:104096. DOI: 10.1016/j.jbi.2022.104096
[16] Zenker S, Strech D, Jahns R, Müller G, Prasser F, Schickhardt C, Schmidt G, Semler SC, Winkler E, Drepper J. National standardisierter Broad Consent in der Praxis: erste Erfahrungen, aktuelle Entwicklungen und kritische Betrachtungen [Nationally standardized broad consent in practice: initial experiences, current developments, and critical assessment]. Bundesgesundheitsblatt Gesundheitsforschung Gesundheitsschutz. 2024 Jun;67(6):637-647. DOI: 10.1007/s00103-024-03878-6
[17] Koll CEM, Hopff SM, Meurers T, Lee CH, Kohls M, Stellbrink C, Thibeault C, Reinke L, Steinbrecher S, Schreiber S, Mitrov L, Frank S, Miljukov O, Erber J, Hellmuth JC, Reese JP, Steinbeis F, Bahmer T, Hagen M, Meybohm P, Hansch S, Vadász I, Krist L, Jiru-Hillmann S, Prasser F, Vehreschild JJ; NAPKON Study Group. Statistical biases due to anonymization evaluated in an open clinical dataset from COVID-19 patients. Sci Data. 2022 Dec 21;9(1):776. DOI: 10.1038/s41597-022-01669-9
[18] ICD-10-GM: Internationale statistische Klassifikation der Krankheiten und verwandter Gesundheitsprobleme. German Modification. [cited 2025 Feb 20]. Available from: https://www.bfarm.de/DE/Kodiersysteme/Klassifikationen/ICD/ICD-10-GM/_node.html
[19] Parker CN, Francis A, Finlayson KJ. Methods for chronic wound research – A scoping systematic review of the recommendations, guidelines and standards. WP&R Journal. 2019;27(2):62-73. DOI: 10.33235/wpr.27.2.0001
[20] Sweeny L. k-anonymity: a model for protecting privacy. International Journal of Uncertainty, Fuzziness and Knowledge-Based Systems. 2002;10(05):557-570.
[21] Ballou DP, Pazer HL. Modeling Data and Process Quality in Multi-Input, Multi-Output Information Systems. Management Science. 1985;31(2):150-162. DOI: 10.1287/MNSC.31.2.150
[22] Wang RY, Strong DM. Beyond Accuracy: What Data Quality Means to Data Consumers. Journal of Management Information Systems. 1996;12(4):5-33. DOI: 10.1080/07421222.1996.11518099
[23] OPS: Operationen- und Prozedurenschlüssel (OPS). [cited 2025 Feb 20]. Available from: https://www.bfarm.de/DE/Kodiersysteme/Klassifikationen/OPS-ICHI/OPS/_node.html
[24] Regenstrief Institute Inc. UCUM. [cited 2025 Feb 20]. Available from: https://ucum.org/
[25] Medizininformatik Initiative – Modul Laborbefund – Implementation Guide. 2021 [cited 2025 Feb 20]. Available from: https://www.medizininformatik-initiative.de/Kerndatensatz/Modul_Laborbefund/IGMIIKDSModulLaborbefund.html
[26] MII IG Fall v2025. 2024 [cited 2025 Jul 22]. Available from: https://www.medizininformatik-initiative.de/Kerndatensatz/KDS_Fall_V2025/MIIIGModulFall.html
[27] Karagounis S, Sarkar IN, Chen ES. Coding Free-Text Chief Complaints from a Health Information Exchange: A Preliminary Study. AMIA Annu Symp Proc. 2021 Jan 25;2020:638-647.
[28] Überleitungsbogen Chronische Wunde 1.0.0. 2023 [cited 2025 Feb 20]. Available from: https://mio.kbv.de/display/UCHW1X0X0